2017 Winner & Scale Up Runner 2018
Real-time diagnostics for devastating wheat rust
The Inspire Challenge is an initiative to challenge partners, universities, and others to use CGIAR data to create innovative pilot projects that will scale. We look for novel approaches that democratize data-driven insights to inform local, national, regional, and global policies and applications in agriculture and food security in real time; helping people–especially smallholder farmers and producers–to lead happier and healthier lives.
This proposal was selected as a 2017 winner, with the team receiving 100,000 USD to put their ideas into practice. The team came runners up for the Scale Up award the following year, receiving an additional USD 125,000 for their outstanding ability to demonstrate the project’s proven viability and potential for impact.


Real-time diagnostics for devastating wheat rust in Ethiopia (MARPLE Diagnostics)
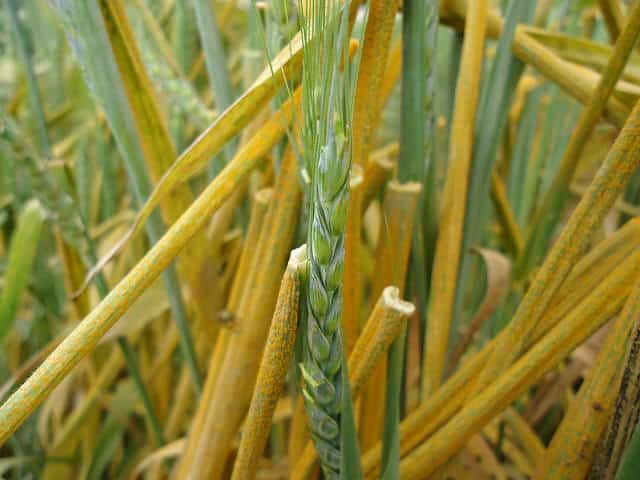
Wheat yellow rust, caused by Puccinia striiformis f. sp. tritici (PST), is considered the most damaging wheat disease globally, causing yield losses sometimes higher than 60 percent. What’s worse is that new PST races have emerged in the last decade that are adapted to warmer temperatures, have expanded virulence profiles, and are more aggressive than previously characterized races. These new strains can lead to large-scale epidemics.
While considerable progress has been made in establishing global wheat rust monitoring systems and surveillance networks, pathogen diagnostics still rely on lengthy and costly controlled bioassays undertaken at a limited number of laboratories.
In order to implement effective early warnings and control measures, new, innovative diagnostic tools are needed to understand the rapidly shifting nature of pathogen populations and dispersal in near real-time.
This project aims to develop and mainstream an affordable, mobile in-field pathogenomics platform called MARPLE (Mobile And Real-time PLant disEase) diagnostics that will revolutionize crop pathogen surveillance and diagnostic. Through quicker, cheaper, and readily deployable software and data management tools, MARPLE diagnostics seeks to reduce wheat yellow rust risks for smallholder farmers.
MARPLE diagnostics will rely on the MinION mobile genome sequencer platform and assess its deployment in Ethiopia.
Project Partners
CIMMYT-Ethiopia is responsible for overall coordination and management of the project and integration of information into the national wheat rust early warning system.
Ethiopian Institute of Agricultural Research (EIAR) is the implementing partner in Ethiopia. EIAR’s National Biotechnology Center at Holeta acts as a centralised hub for the implementation of the MARPLE diagnostics system.
John Innes Centre (JIC) is responsible for the scientific development and testing of the MARPLE diagnostic kit. JIC supports capacity building, logistics, communication activities, and provides technical support to EIAR scientists.
BGRI: Delivering Genetic Gain in Wheat project led by Cornell University financially supports the integration of the MAPRPLE diagnostics system into the established wheat rust early warning system developed by the BGRI programme.

Biotechnology and Biological Science Research Council (BBSRC), UK funded the initial trials of the MARPLE diagnostic system and continue to support JIC staff particularly in training EIAR scientists.
Step by step
US$100K grant
The project was one of five winners of the Inspire Challenge 2017 and was awarded US$100K at the inaugural annual convention of the CGIAR Platform Big Data in Agriculture, 9-22 September 2017.
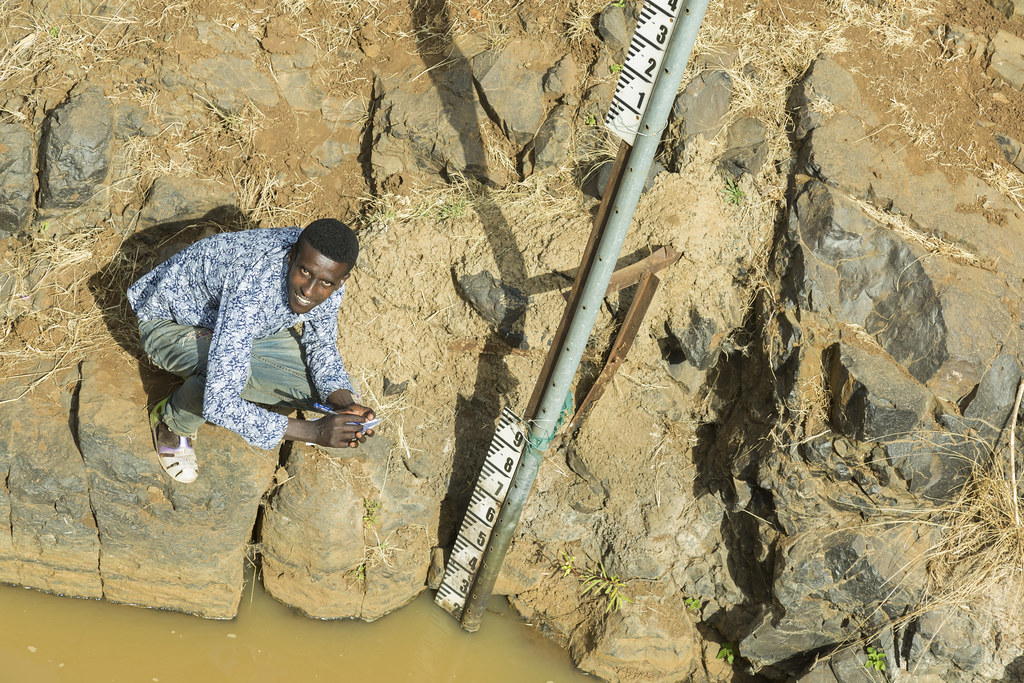
PST sampling

Training of EIAR scientist at JIC, DNA extraction, and creation of sequencing libraries
Dr. Diane Saunders and Abel Mitiku
One Ethiopian scientist was extensively trained at the John Innes Centre (JIC) for five months, whilst others received training directly from JIC scientists in Ethiopia. The team evaluated samples collected in Ethiopia, validated the use of the MARPLE platform, and trained scientists in how to analyze the resulting data in country.
Confirming the accuracy of sequencing on MinION

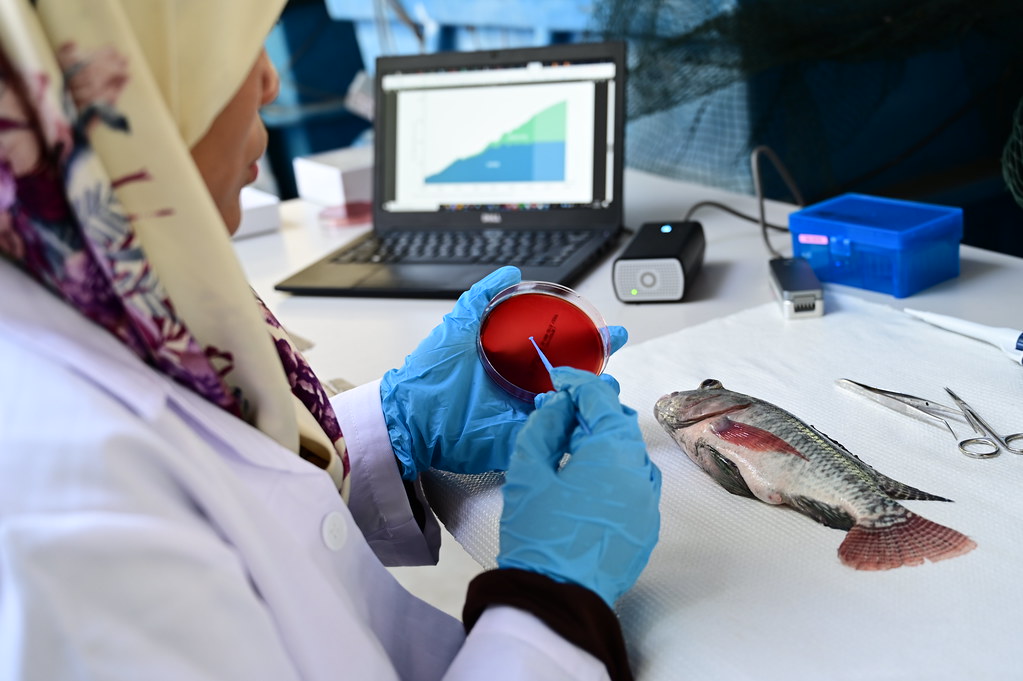
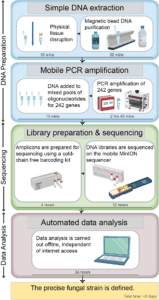
A set of highly variable genes, which discriminated the main genetic groups of yellow rust, were identified from an extensive collection of genomic data from global yellow rust isolates by JIC scientists. Targeted amplicon resequencing of this differential gene panel (approx. 200 genes) using MinION nanopore sequencing and a custom built bioinformatics pipeline then provided an accurate diagnostic.
This scientific methodology, which allows the accurate diagnosis of complex fungal pathogens like PST with large genomes, is a completely new approach.
Piloting MARPLE in Ethiopia
The new and innovative pathogen diagnostics system, which is called MARPLE (Mobile And Real-time PLant disEase diagnostics), was successfully tested in Ethiopia. In the pilot phase, accurate diagnosis of yellow rust strains from field samples to genetic group was obtained within three days of sample collection. This represents a truly significant advance in crop pathogen diagnostics. The pilot demonstrated the first ever accurate identification of individual yellow rust strains whilst crops were at an early growth stage and disease was just starting to emerge in the field. This has never been possible using conventional diagnostics.
Informing disease management decisions in Ethiopia
The data generated by the pilot has been incorporated into national early wheat rust warning systems, illustrating that the MARPLE Diagnostic platform is fully functional and able to generate essential data to inform disease management decisions.
The pilot field testing has rapidly confirmed (in three days) that the PstS11 race of yellow rust is currently present. This new information can help prioritize and target control activities since wheat varieties known to be susceptible can now easily been identified. Successful pilot testing and deployment of the MARPLE diagnostics system in Ethiopia opens the door for technology scaling to help protect wheat crops in East Africa.
US$125K grant
The project was a runner up in the Inspire Challenge Scale Up 2018 and was awarded US$125K at the second annual convention of the CGIAR Platform for Big Data in Agriculture, 3-5 October 2018.
Mainstreaming MARPLE in Ethiopia
This new system is the first operational system in the world using nanopore sequence technology for rapid diagnostics and surveillance of complex fungal pathogens. Furthermore, MARPLE is entirely mobile, deployable in a field-based setting and tailored for use in developing countries. The entire “lab” and analytics fits in a suitcase and runs effectively even in the absence of stable electricity. In addition, the bioinformatics pipeline runs totally offline.
The objective now is to optimize and mainstream the MARPLE diagnostics technique within the wheat research system in Ethiopia. In 2019, the initial focus will be on MARPLE optimization and strengthening a central MARPLE hub. This will lay the foundation to create three additional future MARPLE hubs distributed at key, and strategically located, wheat research stations in the country. This model will ensure that rapid diagnostic capacity for yellow rust is distributed throughout the main wheat growing regions of Ethiopia and will permit sentinel monitoring of yellow rust races appearing on crops on an annual basis.
Expanding to other pathogens
Establishing the distributed MARPLE platform and operating capacity would enable the rapid monitoring system to expand to include additional important pathogens. Additional nanopore sequencing diagnostic methods would need to be developed through future research.
Ethiopia would be the first African nation to have a robust, rapid monitoring system for important transboundary diseases. Given that Ethiopia functions as a gateway for wind-dispersed pests and pathogens between Africa and the Middle East/Asia, the country is the highest priority for such a system.
Co-funding for MARPLE project
Given the successful pilot in Ethiopia, potential for scale-up, and alignment with Delivering Genetic Gain in Wheat (DGGW) activities, co-funding of US$125K was provided to the MARPLE project from the DGGW project led by Cornell University. This support will cover planned MARPLE scale-up activities in 2019.
Developing a road map for application to other important fungal pathogens
A rigorous validation process of the innovative methodology used in MARPLE was undertaken by Diane Saunders group at JIC. A journal manuscript entitled “MARPLE, a point-of-care, strain-level disease diagnostics and surveillance tool for complex fungal pathogens” has been submitted for publication. The article provides a detailed road map for researchers to establish similar systems for other important fungal pathogens.
Rigorous validation
In collaboration with the Global Rust Reference Center and Aarhus University, Denmark, MARPLE sequence results have been aligned with high accuracy to yellow rust reference isolates in current, globally important genetic groups. These genetic groups are the standard way in which yellow rust is being tracked around the world. This alignment will permit standardized reporting of yellow rust genetic groups to the entire global wheat rust community.
Continuous optimization to streamline the MARPLE pipeline
Lab optimization to make MARPLE protocols more efficient and cost effective will be initiated in the Saunders Lab at JIC. Fully optimized, wet-lab protocols using thermostable reagents and library kits that increase sample throughput and run at reduced cost will be developed. The target is to increase sample throughput five-fold and reduce per sample costs by at least 50 percent. Optimized bioinformatics protocols will also be developed.
MARPLE team recognized for international impact
The MARPLE research team — Diane Saunders of the John Innes Centre (JIC), Dave Hodson of the International Maize and Wheat Improvement Center (CIMMYT) and Tadessa Daba of the Ethiopian Institute for Agricultural Research (EIAR) — won the Innovator of the Year 2019 award sponsored by the United Kingdom’s Biotechnology and Biological Sciences Research Council (BBSRC).
BMC Biology paper published
The team published a paper in BMC Biology that shows how the research partnership reduced the speed of diagnostics from many months in high-end labs, to just two days from the side of an Ethiopian field.
The paper outlines the steps that were taken to deliver this combined computational and experimental framework. It is hoped that by publishing this process, similar surveillance methods could be developed for other complex fungal pathogens that pose threats to plant, animal and human health.
Handbook rolled out to MARPLE hubs
A detailed handbook for the MARPLE Diagnostics pipeline will be developed in the Saunders Lab at JIC. This will include all background material required by researchers in Ethiopia to interpret the phylogenetic results.
Establishing additional MARPLE hubs in Ethiopia
Four key wheat research stations will be equipped with MARPLE diagnostic kits in Ethiopia (Kulumsa, Sinana, Mek’ele, and Ambo), and the existing central hub at Holeta will be strengthened.
The Saunders Lab at JIC will arrange procurement and shipping of all components of the MARPLE diagnostic kit that will be distributed to the hubs in-country with assistance from CIMMYT and EIAR.
MARPLE Diagnostics kit demonstrated at meeting with DFID and Gates Foundation
A new partnership between UK aid and the Gates Foundation grants funding to tackle global food insecurity, including funds for scientists researching cutting-edge technology to protect crops.
On 7 October, Bill Gates and Alok Sharma, Secretary of State for International Development of the United Kingdom, visited the Sainsbury Lab at the University of Cambridge. Sharma and Gates participated in a demonstration of the revolutionary MARPLE Diagnostics kit.
Training Ethiopian partners in-country
A comprehensive training course at the Holeta MARPLE hub will be led by Diane Saunders and her team from JIC. Two to four Ethiopian scientists and/or technicians from each of the four hub locations will be fully trained in the entire MARPLE pipeline from sample processing to final data analysis.
2020 will focus on strengthening the established MARPLE hubs across Ethiopia and continuous review and optimization of the methodology.
Four UK PhD students will be stationed at research stations (two at Holeta, two at Kulumsa) to support research activities and training.
Project News and Resources
Meet all the Winners