Agricultural Ontologies in Use: The MIAPPE Standard
The Ontologies Community of Practice is engaged in the development of ontologies for agricultural research. In a series of blog posts, we’ll take a look at ongoing ontologies projects and developments.
MIAPPE was originally developed within the transPLANT project and published in 2015, with the first update, dubbed version 1.1 released in 2019. This update was led by ELIXIR, the European infrastructure for data in life sciences, with support from the international plant sciences community.The goal of the update was to refine and extend the standard in view of the FAIR data principles, to broaden the scope of experiments MIAPPE can support, and to increase its usability and documentation. The input from the plant sciences community, which was instrumental in ensuring that MIAPPE served their needs, was received through both online consultation and in-person events such as the 2018 PhenoHarmonIS workshop.
The MIAPPE reporting standard is organized into 11 sections which span attributes ranging from general research organization details (project, parties responsible), to plant materials and their provenance, experimental set-ups (time, place, purpose, layouts, independent variables), to phenotyping and environmental variables measured. As of version 1.1, MIAPPE can be used for greenhouse, field or in situ experiments about crop plants, model organisms, or woody plants, covering a variety of experimental designs. These use-cases have been validated through implementing MIAPPE 1.1 for datasets representing all these experimental purposes and settings.
MIAPPE 1.1 explicitly declares the relationships between its sections and lists the mandatory and recommended attributes for each of them, as well as their restrictions and requirements. To comply with the FAIR data principles, MIAPPE also includes recommended ontologies for all fields for which adequate ontologies exist, such as the species-specific trait ontologies of the Crop Ontology to describe plant traits and methods to assess them, the Crop Research Ontology to describe experimental design details, and the Plant Ontology to describe plant anatomy and development stages. Furthermore, the use of unique identifiers is recommended for all key concepts throughout the standard.
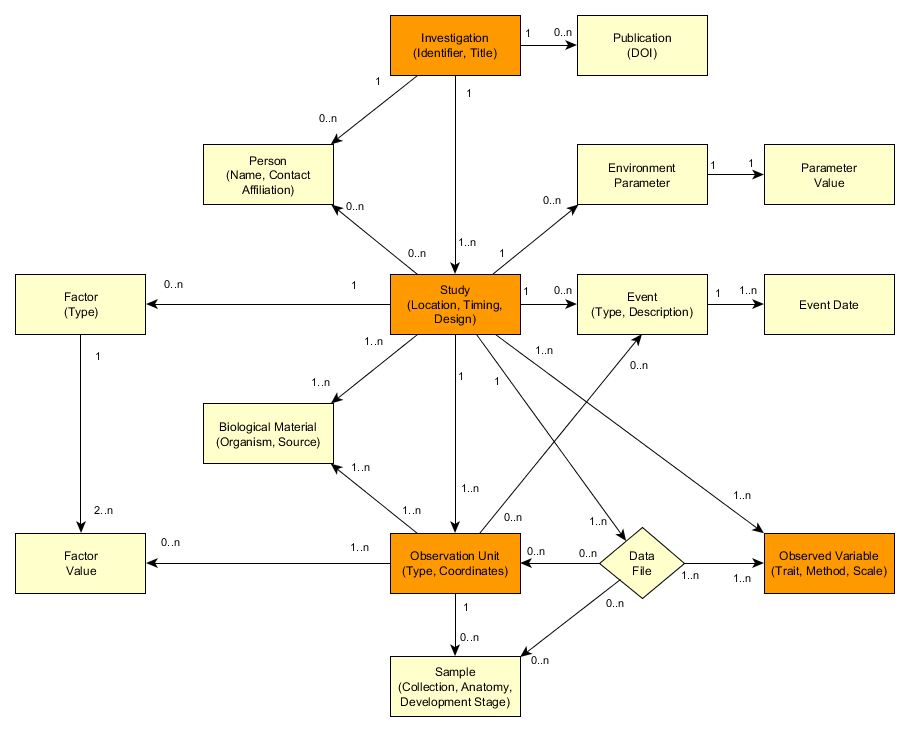
The data model of MIAPPE version 1.1 (simplified).
In addition to its human-readable release, the MIAPPE 1.1 standard was formalized in Web Ontology Language (OWL) as the Plant Phenotype Experiment Ontology (PPEO). Semantic representation unlocks the many benefits of a machine-readable format, such as automated validation of MIAPPE compliance, machine interpretation of MIAPPE-compliant datasets, and interoperability between MIAPPE and other related standards such as the Breeding API (BrAPI).
In PPEO, MIAPPE sections are represented as classes, and MIAPPE attributes are mainly represented as data properties connected to the corresponding classes. Intermediate classes and object properties are used to capture paired or grouped attributes, as necessary. Ontology recommendations for MIAPPE fields are encoded as annotations of the properties or property restriction axioms of classes. Correspondences between MIAPPE fields and BrAPI fields are encoded as cross-references.
The PPEO encoding of MIAPPE enables the publication of MIAPPE datasets in Resource Description Framework (RDF), to complement the MIAPPE implementation in ISA-Tab that was updated for version 1.1 and enriched with additional examples. Furthermore, MIAPPE 1.1 is also implemented in a spreadsheet template, developed mainly for training purposes, where each section of the standard, with its fields, definitions, and examples, comprises one sheet.

Photo: Sureshlm [CC BY 4.0]
Like any standard, MIAPPE is an ongoing effort that must keep up with advances in both plant science and data management practices and technologies, as well as cater to the needs of the community it aims to serve. To better serve the community, we are in the process of developing a web-based graphical user interface with step-by-step instructions for MIAPPE dataset submission. With respect to data management technologies, we are encoding MIAPPE in JSON-schema to support publication of MIAPPE datasets in JSON. Concerning advances in plant science, plans are already in place to tackle the description of sensors and the provision of better guidelines for environmental characterization.
Click here to access MIAPPE’s paper, “Enabling reusability of plant phenomic datasets with MIAPPE 1.1.”
January 27, 2020
Eliana Papoutsoglou on behalf of The MIAPPE Team
Latest news